Discover Our Latest Research
Explore groundbreaking studies and find inspiration.

119.
Split-APEX implicates splicing factor SRSF1 and splicing helicases in ribosomal biogenesis
118.
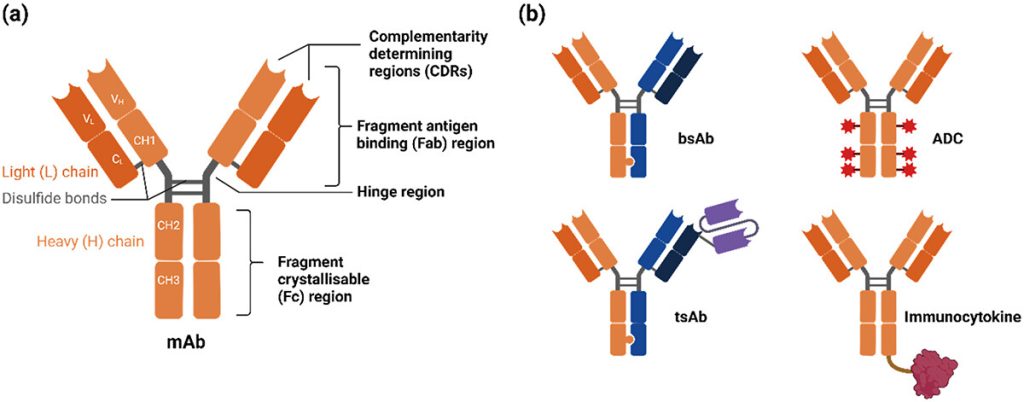
Antibody-drug conjugates as multimodal therapies against hard-to-treat cancers
117.
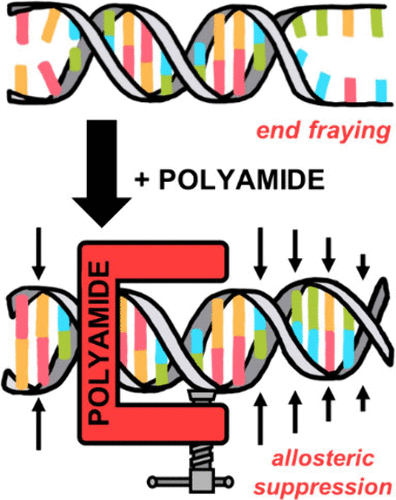
Long-Range Allosteric Modulation of DNA Duplex Dynamics Induced by Pyrrole-Imidazole Polyamide Binding
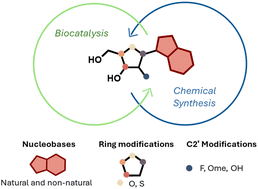
115.
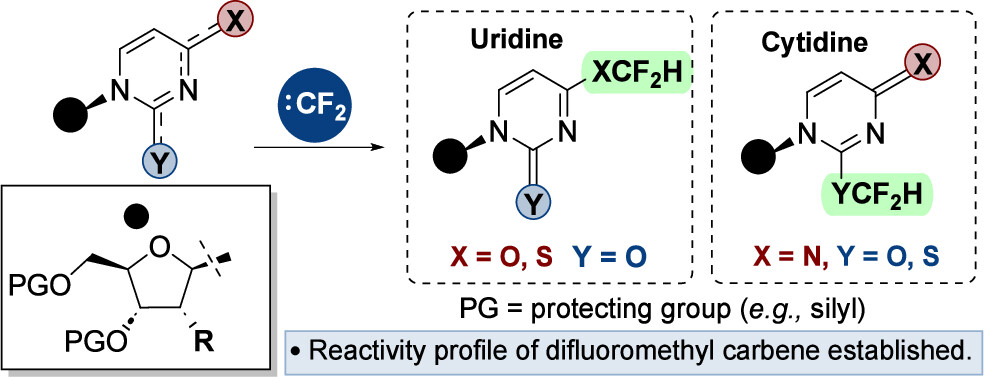
Preparation of nucleoside analogues: opportunities for innovation at the interface of synthetic chemistry and biocatalysis

113.
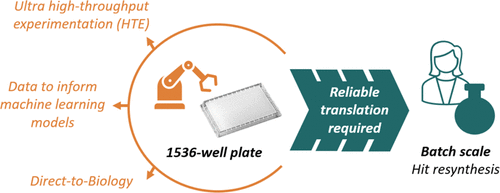
Factors to Consider for Synthesis in 1536-Well Plates─An Amide Coupling Case Study for PROTAC Synthesis
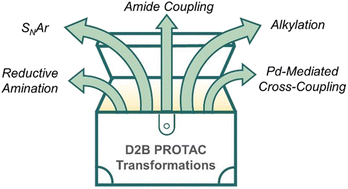
112.
Expanding the reaction toolbox for nanoscale direct-to-biology PROTAC synthesis and biological evaluation
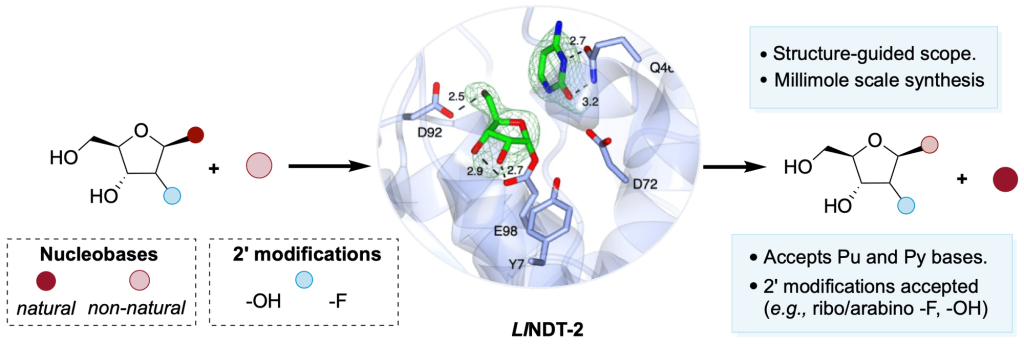
111.
Biocatalytic Synthesis of Ribonucleoside Analogues Using Nucleoside Transglycosylase-2
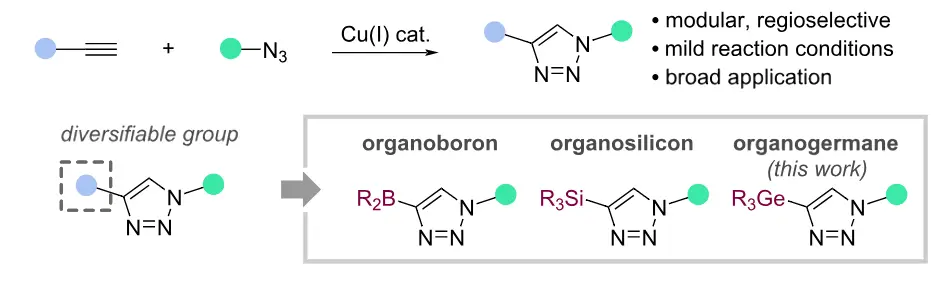
110.
Germanyl triazoles as a platform for CuAAC diversification and chemoselective orthogonal cross-coupling
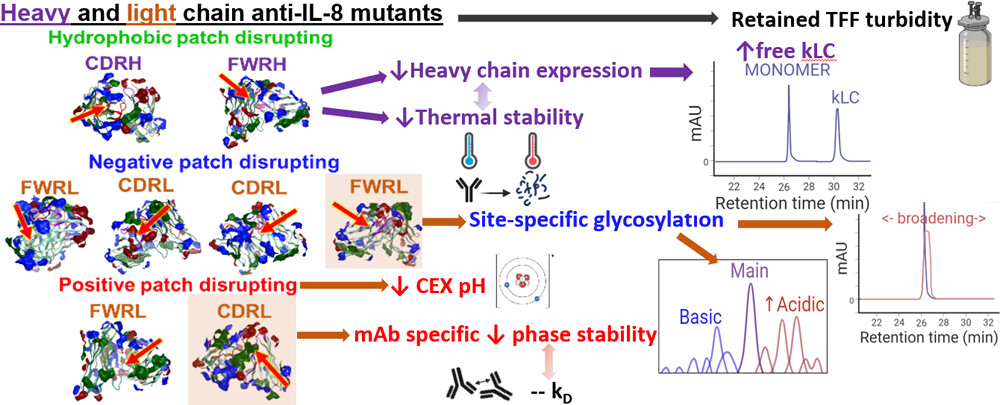
109.
Assessing the Manufacturability and Critical Quality Attribute Profiles of Anti-IL-8 Immunoglobulin G Mutant Variants

108.
Core Modifications of GSK3335103 toward Orally Bioavailable αvβ6 Inhibitors with Improved Synthetic Tractability
Let's Partner for Progress
We invite researchers and institutions to join us in advancing science. Together, we can exchange ideas and drive innovative research in chemistry.